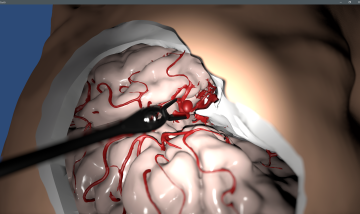
Artificial intelligence as the key to recognizing brain arteries
Artificial intelligence is opening up new avenues in medical imaging: advanced algorithms can now also identify and annotate cerebral arteries in non-angiographic images, enabling a broader diagnosis and treatment of cerebral vascular diseases.
by Bertram Sabrowsky-Hirsch, MSc
Contents
- Innovation in medical imaging: AI discovers brain arteries
- Decoding the Circle of Willis: Preventing neurological crises through imaging
- Insight into the tissue jungle: MR imaging reveals blood vessels
- Methodology of the future: AI-supported segmentation of brain arteries
- Results of innovative vessel segmentation in MR sequences using AI methodology
- Conclusion: Breakthrough in the annotation of cerebral arteries with AI
- Acknowledgments
- References
- Author

Innovation in image-based diagnosis: AI discovers brain arteries
Imagine if technology could detect the risk of stroke before it’s too late. Stroke is the second leading cause of death worldwide, claiming countless lives and often leaving survivors with severe neurological damage [1]. In Germany, one in 40 adults has already had a stroke [2]. This is where neurological imaging plays a critical role – it is the key to diagnosing and treating these devastating diseases.
But the challenge: blood vessels are difficult to recognize on conventional images. Traditionally, medicine has relied on angiographic methods, which are effective but not always available and often require the use of contrast agents. This is not only costly, but can also pose risks for patients.
This is where the groundbreaking work of RISC Software GmbH comes in. In the publication “Segmentation and Anatomical Annotation of Cerebral Arteries in Non-Angiographic MRI”, the researchers explore how artificial intelligence can help visualize blood vessels in standard MR images – a revolution in imaging that paves the way for earlier and broader diagnoses. This innovative method, which also works without special angiographic image data, opens up a new field for concomitant diagnoses and makes it possible to re-evaluate historical data. In the future, AI could help to prevent strokes.
Examination of the Circle of Willis: Prevention of neurological emergencies through imaging
Directly below the brain is a vital arterial structure known as the Circle of Willis (CoW). Its unrestricted function is essential for the prevention of neurological emergencies, as disruptions can lead to serious health consequences. Modern imaging techniques open up new possibilities to assess the hidden aspects of this central vascular structure, enabling early detection and prevention of neurological events.
High-resolution scans allow physicians to view the complex branches and connections of the CoW in detail and detect pathological conditions such as aneurysms, stenoses, occlusions and vascular malformations that are more common in this area. These valuable insights into the cerebral circulation are key to preventing strokes and other life-threatening complications by enabling timely therapeutic interventions and preserving patient health.
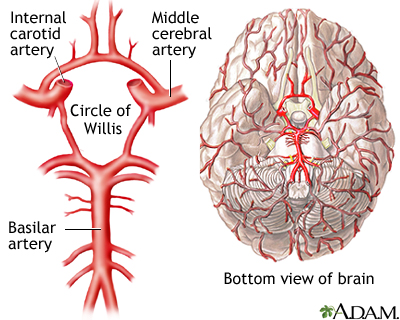
Figure 1: The arterial ring of the Circle of Willis supplies the brain with blood.
(Source: MedlinePlus, https://medlineplus.gov/ency/imagepages/18009.htm)
Insight into the tissue jungle: MR imaging reveals blood vessels
How easily blood vessels can be differentiated from the surrounding structures in the image data depends primarily on the imaging method used. The advantage of magnetic resonance (MR) imaging is that it is non-invasive and does not require the use of X-rays. In addition, radiologists have different sequences at their disposal to emphasize different tissue properties and thus highlight certain anatomical structures. The time-of-flight (TOF) sequence, for example, enables the blood flow within the arteries to be imaged, but is less suitable for assessing brain tissue than the T2 sequence due to its low contrast. Which sequences are used for image acquisition depends on the medical indication and the number of different sequences is limited in favor of the acquisition time. Image data is therefore often not available, which makes it difficult or even impossible to assess the blood vessels. Figure 2 illustrates the visibility of blood vessels in different MR sequences.
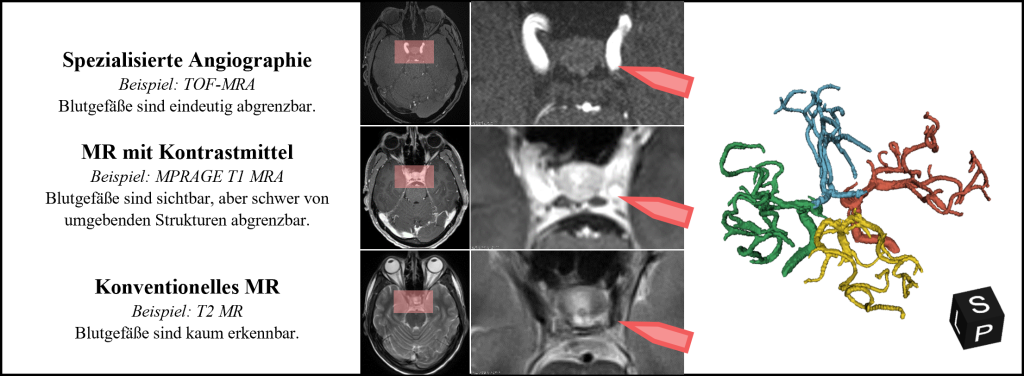
Figure 2: The visibility of blood vessels strongly depends on the imaging method used. In the sectional images, the same section of a cerebral vascular tree (right) is shown in three different sequences.
Methodology of the future: AI-supported segmentation of brain arteries
Researchers at the Department of Medical Informatics have developed a fully automated method that segments the cerebral arteries along the Circle of Willis and annotates four vascular regions (anterior, posterior, left and right). Their method supports three different non-angiographic MR sequences as well as the angiographic TOF-MRA sequence. For this purpose, an artificial intelligence was trained on 555 patients from the public IXI dataset [3]. The MR sequences PD, T1, T2 and TOF-MRA are available for each patient in this dataset. The segmentation task for the angiographic TOF-MRA image data could be solved automatically using traditional image processing methods (see Figure 3), and the annotations were then successfully transferred to the non-angiographic sequences (see Figure 4). The artificial intelligence was then trained to master the segmentation task for all available sequences. Specifically, semantic segmentation models based on artificial neural networks (ANN) were developed using the nnU-Net framework [4].
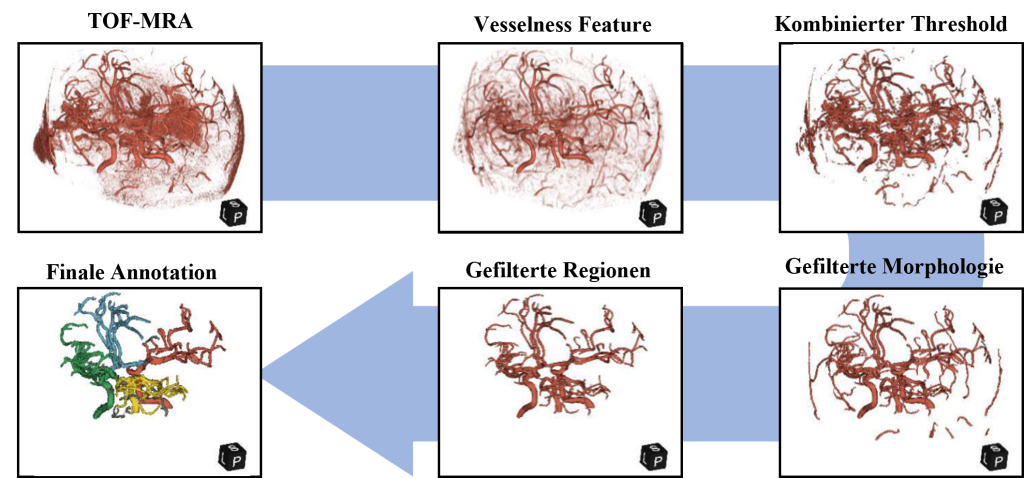

Figure 3: The automatic method for annotating the blood vessels and vessel regions in the angiographic TOF-MRA sequence is based on traditional image processing methods.
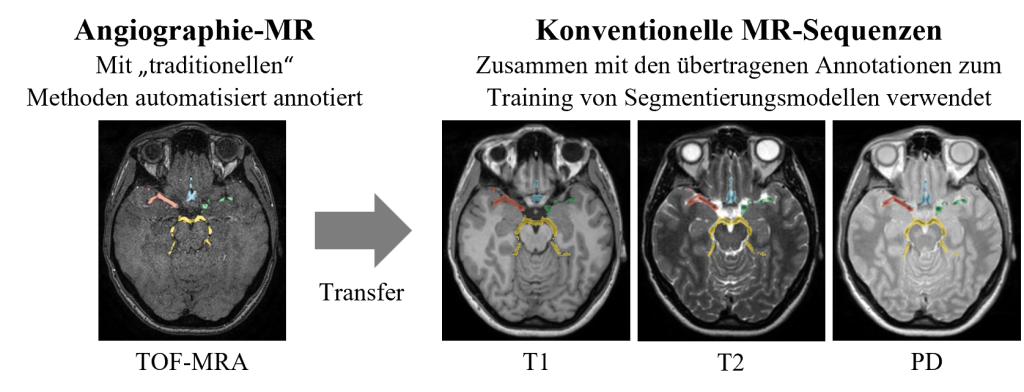
Figure 4: The annotated blood vessels from the TOF-MRA sequence were transferred to the non-angiographic sequences (PD, T1, T2). The resulting dataset of image data and ground truth annotations was then used to train the segmentation models.
Results of innovative vessel segmentation in MR sequences using AI methodology
The researchers from the Department of Medical Informatics carried out the training and evaluation of their method using 5-fold cross-validation on 485 patients.inside. In addition, 70 of the original 555 patients were The test data set was analyzed using a different imaging hardware at a different institute. This strategy was used to test the applicability of the method across different institutes.
The method was evaluated by the researchers through visual inspection and quantitative analysis of similarity measures based on the generated annotations. For this purpose, established metrics such as the Dice Similarity Coefficient (DSC), the 95 percentile of the Hausdorff distance (HD95) and the average symmetrical surface distance (ASD) were calculated. In the cross-validation, the DSC around the Circle of Willis achieved a very good value of 0.82 (in the literature, a value of 0.7 is considered to be of high quality [5]). The ASD metric showed a deviation of 0.44 mm, which is a very good result considering the average resolution of the image data of 0.92 mm. The results indicate that vessel structures in the MR sequences examined can be segmented and annotated with adequate quality.
For the additional test data set, the models achieved a DSC of 0.75, which is still a good result despite the significant reduction. However, this indicates that the method should be improved by training with more extensive data sets from different institutes in order to achieve more robust performance with different imaging hardware.
There were clear differences between the individual MR sequences: Although the angiographic TOF-MRA sequence provided the best results, and the segmentation task was also relatively easy here due to the clear visibility of the blood vessels. Based on the results, the PD sequence (weighted by proton density) was best suited for annotation, followed by T2 and T1. Small differences were also observed with respect to the vascular regions, with better results for the left and right regions than for the anterior and posterior regions.
Comparable studies based on conventional, non-angiographic image data are lacking in the literature. Klimont et al [6] described a method for non-angiographic CT image data, but not for MR sequences. The researchers’ results for the TOF-MRA sequence are comparable to values from the literature [7] and offer a unique advantage by supporting non-angiographic MR sequences.
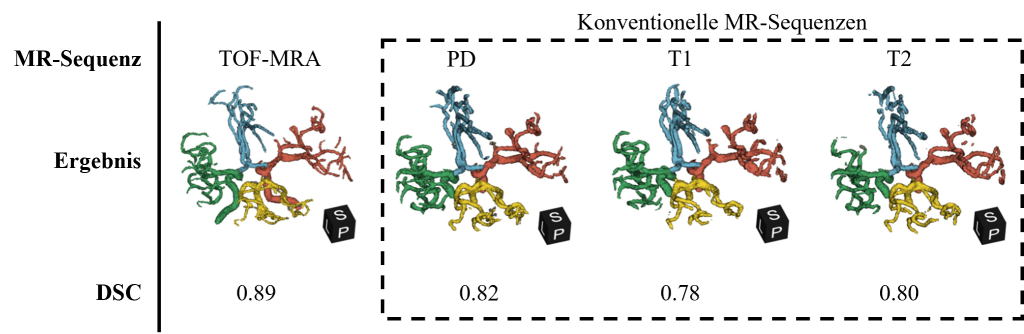
Figure 5: Visual representation of the results using a patient from the cross-validation data compared to the different MR sequences. The DSC metric refers to the average value in the Circle of Willis area for the respective sequence from the cross-validation.
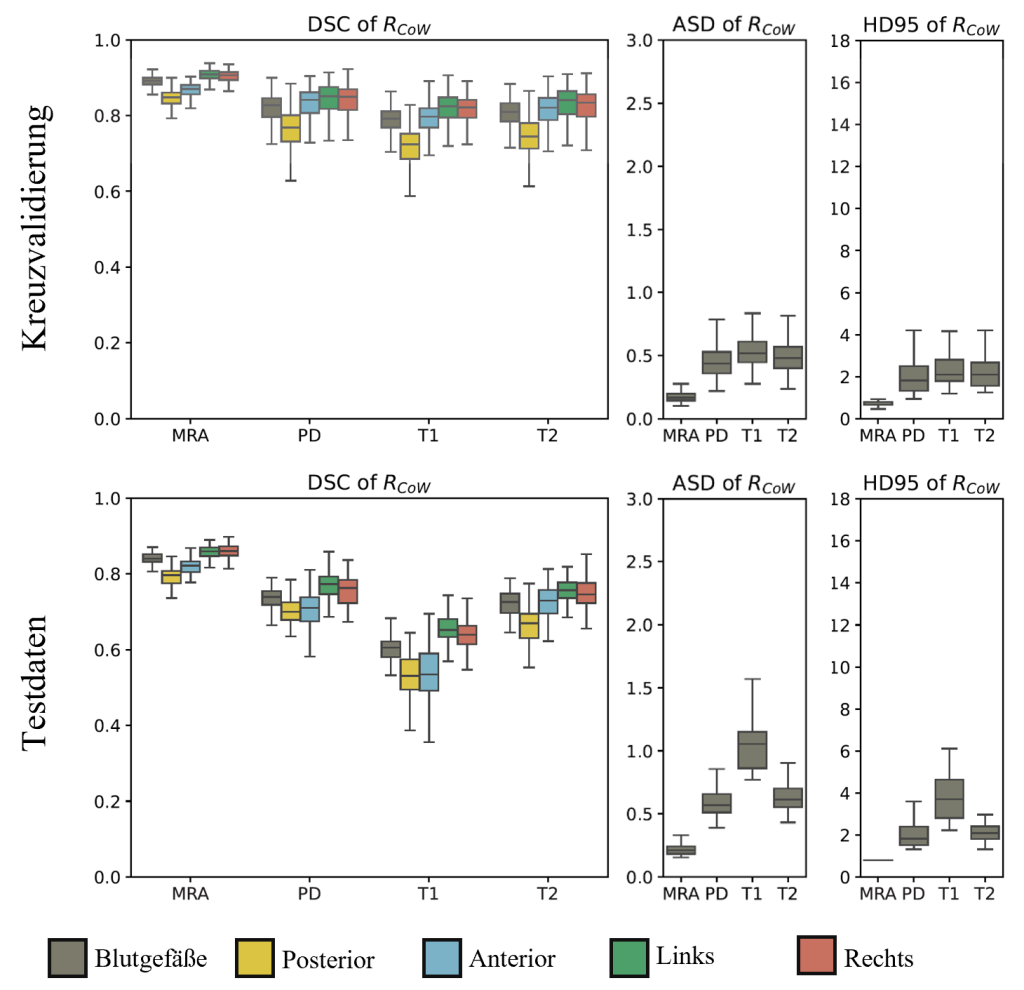
Figure 6: Detailed evaluation of the DSC, ASD and HD95 metrics in the Circle of Willis area as part of the cross-validation (top) and for the additional test data (bottom). The blood vessels were evaluated in a general label and additionally for each vessel region (posterior, anterior, left, right). The results refer to the respective MR sequence: (TOF-)MRA, PD, T1 and T2.
Conclusion: Breakthrough in the annotation of cerebral arteries with AI
In their experiments, researchers from the Department of Medical Informatics have demonstrated that artificial intelligence can also be used to annotate blood vessels in non-angiographic image data with high quality. This represents a significant advance in the field of analyzing medical image data, as previous methods focused mainly on angiographic image data. For a broader applicability of the method, an extension of the training data is necessary both in scope and in variability with regard to imaging hardware and medical indications. In particular, the inclusion of data from patients with aneurysms and tumors is crucial for a robust segmentation of these malformations. The researchers are currently continuing to work on supporting additional MR sequences such as the MPRAGE MRA sequence.
At present, the researchers recommend using their method primarily to localize the Circle of Willis and not to assess the vascular structures, as the correct annotation of malformations cannot yet be verified. The researchers are currently using the annotations created to localize and automatically register the blood vessels with DSA (Digital Subtracted Angiography) image data. However, planned extensions could make the method usable for the analysis of blood vessels in non-angiographic image data in the future.
The results and methodology were presented at the 2023 DMIP conference in Japan and are available in the Proceedings of the ACM Digital Library (ISBN: 979-8-4007-0942-5). The annotated dataset and the trained models are publicly available in the department’s GitHub repository.
Link to the repository: https://github.com/risc-mi/cerebral-artery-annotation.
Acknowledgments
This work was funded by the FFG (Austrian Research Promotion Agency) as part of Grant 872604 (MEDUSA) and by research funding from the Province of Upper Austria. RISC Software GmbH is part of the UAR (Upper Austrian Research) network.
References
[1] Katan, M. and Luft, A. (2018). Global Burden of Stroke. Semin Neurol, 38(2), 208-211. https://doi.org/10.1055/s-0038-1649503
[2] Robert Koch Institute (ed.) (2015) Health in Germany. Federal health reporting. Jointly supported by RKI and Destatis. RKI, Berlin, https://doi.org/10.17886/rkipubl-2015-003-2
[3] Information eXtraction from Images (IXI) Dataset. (Retrieved March 23, 2023) from https://brain-development.org/ixi-dataset/
[4] Isensee, F., Jaeger, P. F., Kohl, S. A. A., Petersen, J., & Maier-Hein, K. H. (2021). nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods, 18(2), 203-211. https://doi.org/10.1038/s41592-020-01008-z
[5] Zou, K. H., Warfield, S. K., Bharatha, A., et al. (2004). Statistical validation of image segmentation quality based on a spatial overlap index. Acad Radiol, 11(2), 178-189. https://doi.org/10.1016/s1076-6332(03)00671-8
[6] Klimont, M., et al. (2020). Deep learning for cerebral angiography segmentation from non-contrast computed tomography. PLOS ONE, 15(7), e0237092. https://doi.org/10.1371/journal.pone.0237092
[7] Hilbert, A., et al. (2020). BRAVE-NET: Fully Automated Arterial Brain Vessel Segmentation in Patients With Cerebrovascular Disease. Frontiers in Artificial Intelligence, 3rd ed. https://doi.org/10.3389/frai.2020.552258
Ansprechperson
Author

Bertram Sabrowsky-Hirsch, MSc
Researcher & Developer